The following steps need to be done for getting the gm/Id vs Id/(W/L) graphs in Cadence for a MOS transistor.
Create the schematic
The schematic is a simple transistor schematic. The schematic below shows an example: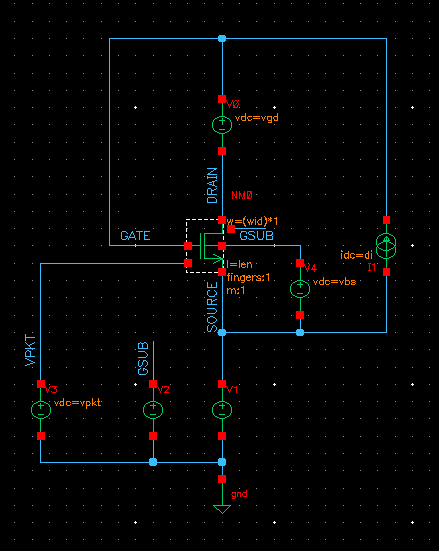
The above schematic shows that the drain current(di), vgd, vbs, W and L are parameterized and they are sweeped in the simulation script (except vbs) to get the graphs for a wide range of conditions and check the variation of the graph for them.
Save Operating Point Info
Create a text file called mysaves_t18.scs and add the following line in it:Now add this file with path in ADE->Setup->Simulation Files->Definition Files
Note that the .m0 is only needed because this model is implemented as s Sub Circuit in the models file. If the model is not a subcircuit and something direct like a BSIM model then .m0 would not be needed.
Setup simulation in ADE
Setup the DC sweep simulation in ADE by DC sweeping di from a low value to a high value for a typical vgd, W and L. Set the sweep to be logarithmic with 50 points per decade. Also setup expressions to calculate- gm/Id = gm/di expression: (getData("MNO_PSP.m0:gm" ?result "dc-dc")/IS("/V1/PLUS"))
- Id/(W/L) expression: (IS("/V1/PLUS")/(5e-6/180e-9))
Run the Simulation and see the results.
Create the Ocean Script
Save the Ocean Script from the ADE environment and modify it to look like the following:Save this ocean script (say 'gmByIdScript.ocn') and load it from the CIW window by typing:
This would start the simulations and at the end we would have the file gmByIdCurves.csv with all the simulation results.
Draw the Graph using MATLAB
Finally create the following script in MATLAB and name it mergeGraphs.m and run it to create the graphs in a figure:Drawing the Graph using gsl-shell
Use the following script. The script can be loaded in gsl-shell using the command dofile('script_path_and_name').Sample Graph
Shown below is a sample graph from the above steps: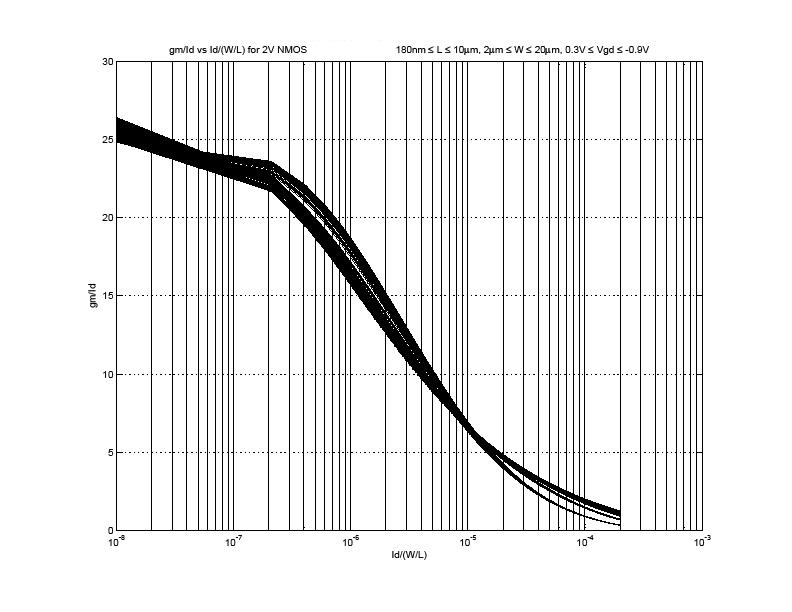
NOTE This graph was taken with a linear sweep a better graph will result of the sweep is set to logarithmic especially with no discontinuity in the slope as it seems in this one.